Deep learning software tools to the rescue of medical research
Jaspreet Bindra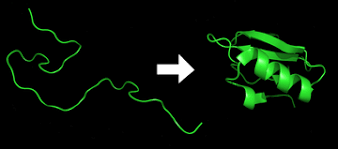
Move Thirty-Seven, five lines from the edge of the board, was a blinder. Startling and totally unexpected, it was proclaimed as a ‘clear mistake’ by watching experts. Lee Sedol, the opponent, flinched and was forced to think for twelve minutes before he could make his rejoinder. One expert, Fan Hui, watching from the sidelines saw through it, however: “It’s not a human move. I have never seen a human play this move. So beautiful. Beautiful. Beautiful. Beautiful.” It was 2016, the game was Go, and the undefeated world champion Sedol was playing with a machine, AlphaGo, created by an Artificial Intelligence company called DeepMind. AlphaGo had defeated Lee in the first game, this ‘inhuman’ move steered it to a two-zero lead. This was a Sputnik moment in AI, since Go, unlike chess, is an intricate, non-deterministic game. A game of instinct and feeling, it has ten to the power 170 possible moves, more moves than the ‘atoms in the known universe’.
The force that AlphaGo unleashed is the power of Deep Learning, a subfield of machine learning built with algorithms inspired by the structure and function of the brain, also known as neural networks. DeepMind, the pioneer of this field, has used it to power face-recognition cameras and voice assistants and defeat humans in multiple games. But its most astonishing application was unveiled last week – predicting how proteins fold.
“To understand life”, says The Economist, “you must understand proteins. These molecular chains, each assembled from a menu of twenty types of amino acids, do biology’s heavy lifting. In the guise of enzymes, they catalyze the chemistry that keeps bodies running. Actin and Myosin, the proteins of muscles, permit those bodies to move around. Keratin provides their skin and hair. Haemoglobin carries their oxygen. Insulin regulates their metabolism. And a protein called spike allows coronaviruses to invade human cells, thereby shutting down entire economies.” Proteins are the origin of existence; the tail of a human sperm is a structure composed of many types of proteins that work together to form a complex rotary engine that propels the sperm forward to fertilize an egg, and to create life.
Proteins also do something else: they fold. The final, intricate shape they take after folding determines their function. For instance, one of them can fold like “snakes in a can”; which when embedded in a cell membrane, creates a tunnel that allows traffic into and out of cells. Other proteins form shapes with pockets called “active sites” that are perfectly shaped to bind to a particular molecule, like a lock and key, the spike on the coronavirus. By folding into myriad shapes, proteins perform different roles despite being composed of the same basic building blocks. The Harvard Business Review draws an analogy: |all vehicles are made from steel, but a racecar’s sleek shape wins races, while a bus, dump truck, crane, or excavator are each shaped to perform their own unique tasks”. If they fold wrongly, as they often do, they can cause horrific harm; the accumulation of misshapen proteins is said to cause Alzheimer’s, Parkinson’s, Huntington’s, and Lou Gehrig’s (ALS) disease. Wrong folds also cause cystic fibrosis and sickle cell anemia, and thousands of diseases we do not know yet.
Therefore, predicting how a protein will fold has been one of the biggest and most difficult challenges in science for decades. Scientists have used techniques like x-ray crystallography, but these are slow and usually after the fact. To solve this humongous challenge is where DeepMind’s learnings from AlphaGo came in to create AlphaFold 2, a deep learning program to predict how a protein will fold. Proteins are even more complex than Go; a protein could take any of as many as ten raised to the power 300 different shapes. Go players do not play the game by knowing every step, or by brute force as chess can be, but by ‘taking short-cuts’, by intuition and by what ‘feels right’. This is exactly how people played a game called FoldIt, which was a simulation on protein folding. DeepMind saw this analogy and used the same principles it used to create AlphaGo for AlphaFold. By feeding powerful computers earlier examples and patterns, they taught them learn to apply these intuitive short cuts and rule-of thumbs, similar to what humans would do, but strain to articulate. Much like the aforementioned Move Thirty-Seven, the computers would sometimes come up with solutions and insights that would stun human experts. In the latest test, a competition called CASP, AlphaFold 2 got a score of 92.4, way above anything ever before, handing a powerful tool in the hands of medical scientists. Knowing how proteins fold can help us address so far incurable diseases in a totally new way, and also prevent many from happening. As Andrei Lupas of Max Planck Institute exclaimed, “This will change medicine. It will change research. It will change bioengineering. It will change everything.” One of the greatest mysteries in science has been unfolded.
(This article was first published as an OpEd in Mint dated Dec 10, 2020)


